| The
POOL Lab Research People Publications Genomes |
|
Why study Population Genetics?
by John Pool Historical perspective The origins of population genetics are inextricably tied to the modern synthesis of biology, combining the great insights of Darwin, Mendel, and their intellectual descendents to provide a concrete framework for how evolutionary forces like natural selection and genetic drift can act upon heritable traits. As an undergraduate, one of the most appealing things to me about population genetics was how it connected my interests in genetics and molecular biology on one hand with evolution and ecology on the other, and I still appreciate having research that interfaces with such diverse aspects of biology. The University of Wisconsin has had a critical role in the development of this field. One of the primary architects of population genetics, Sewall Wright, spent the latter part of his career here (1955 to 1988). Wright was drawn by the presence of James Crow, who had a long and very productive career at UW Madison from 1948 to 2012. Crow, perhaps more than anyone else, helped to bridge classical and modern population genetics (he interacted with founders of the field, and ultimately trained many of today's prominent population geneticists). He was also known for helping his Ph.D. student Motoo Kimura to develop the influential neutral theory of molecular evolution. I was fortunate enough to overlap here in Madison with this greatly admired and universally liked scientist, if only briefly, and he was still a regular on campus until his passing at the age of 95. In its early years, population genetics was largely a theoretical discipline - constructing elegant models of how genetic variation should look in nature under various evolutionary models, but in those days genetic variation could almost never be observed. Even within the field, many once viewed population genetics as an abstract disclipine with limited practical application. The genomic revolution Theoretical advances continue to play a meaningful role in modern population genetics, but in recent years our ability to generate data that directly measures genetic variation has increased by orders of magnitude. In 2007, genetic variation was named by the journal Science as "breakthrough of the year", and today we can sequence large numbers of genomes for organisms ranging from Drosophila to our own species. Genomics has touched nearly every area of biology, but for population genetics, it has truly ushered in a golden age for the field. Today we can examine genetic variation at any gene in the genome, conduct comprehensive genomic scans for targets of recent natural selection, obtain more accurate estimates of population history, and perhaps generate enough data to test the population genetic models that have been developed and debated over several decades. 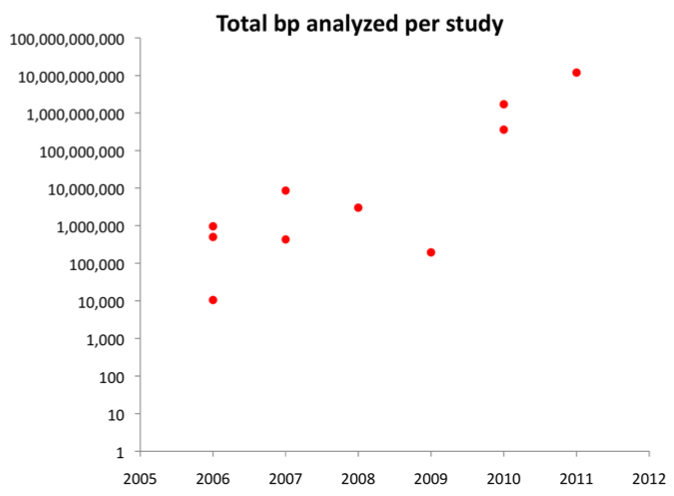 To illustrate how fast population genetic data sets are growing, here is a log-scale plot showing the total number of DNA bases analyzed in my first 10 publications (arranged by year). As a grad student, I started out sequencing one gene at a time in small sets of Drosophila fly stocks. Today my lab analyzes large numbers of fully sequenced genomes. Ultimately our improved ability to study genetic variation will have wide-ranging practical implications as well, for fields ranging from agriculture to conservation to medicine. More than one population geneticist, after spending most of a career on purely academic pursuits, now finds themself consulted by pharmaceutical companies for advice or collaboration, as the promise of personalized medicine informed by individual genotype moves closer to reality. Of course, there is still much work to be done. We have really just collected the first few population genomic data sets and performed the first few statistical analyses on these data. Many of the field's great mysteries remain unresolved, like the importance of natural selection in shaping genetic variation, and the types of genetic changes that contribute to adaptive evolution. It's an excellent time for collecting new data and for developing new ways of analyzing it. Outlook for young researchers in population genetics The academic job market is always tough, but relative to other fields that I'm familiar with, I've seen a lot of my peers in population genetics land strong faculty positions. Interest from the private sector may increase as well, as genetic variation becomes a more immediate concern for tasks ranging from drug design to biofuels and crop improvement. In general, researchers with experience collecting and especially analyzing genome-scale data have a big advantage finding jobs after grad school. For undergrads still exploring their interests, biological knowledge and lab experience are still important, but having a little background in computer programming and statistics can really help too (whether you go into population genetics or not). This doesn't mean you should be dissuaded if you don't have a lot of math and computer science courses as an undergrad (I didn't) - just be ready to learn some new skills as you go along (I certainly still do). |
Research People Publications Why Study Population Genetics? |